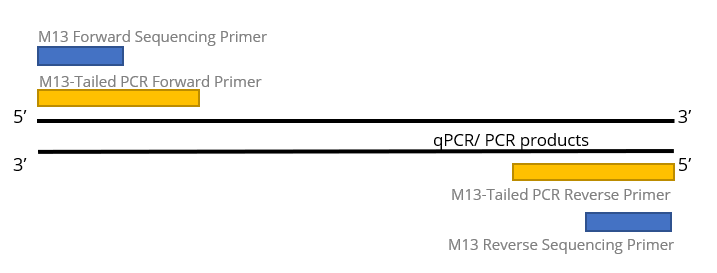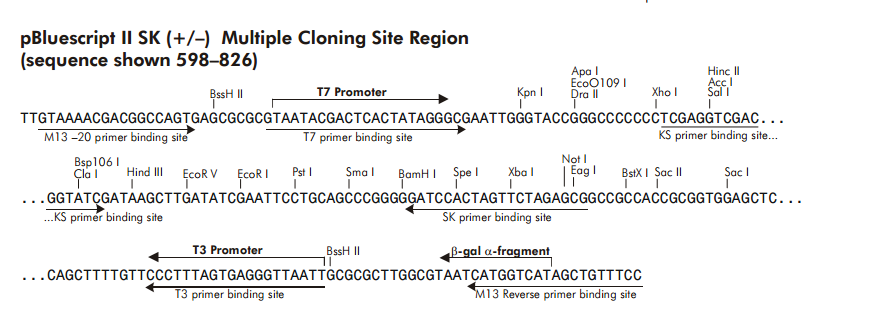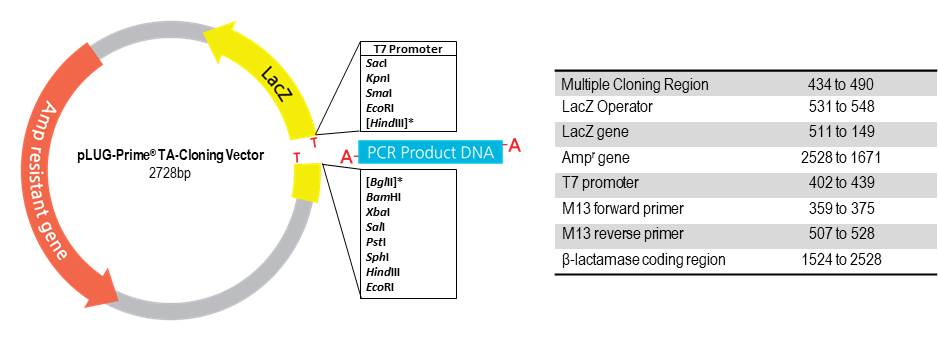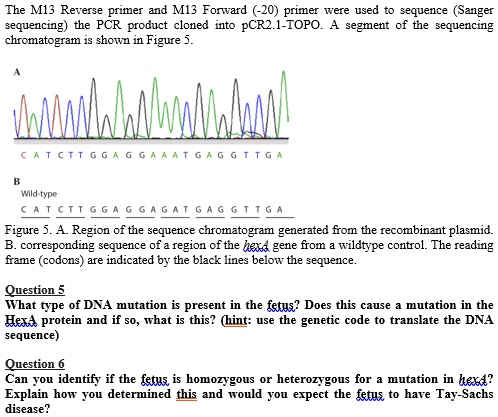
SOLVED: The M13 Reverse primer and M13 Forward (-20) primer were used to sequence (Sanger sequencing) the PCR product cloned into pCR2.1-TOPO. A segment of the sequencing chromatogram is shown in Figure
M13-Tailed Simple Sequence Repeat (SSR) Markers in Studies of Genetic Diversity and Population Structure of Common Oat Germplasm | SpringerLink

PacBio on X: "A simple way to segue from Sanger to #SMRT Sequencing in the clinical lab? The answer lies in barcoding, and we teamed up with @LABCORP and @SageSci to test

An Efficient Technique for Primer Development and Application that Integrates Fluorescent Labeling and Multiplex PCR
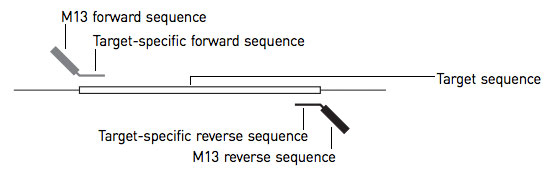
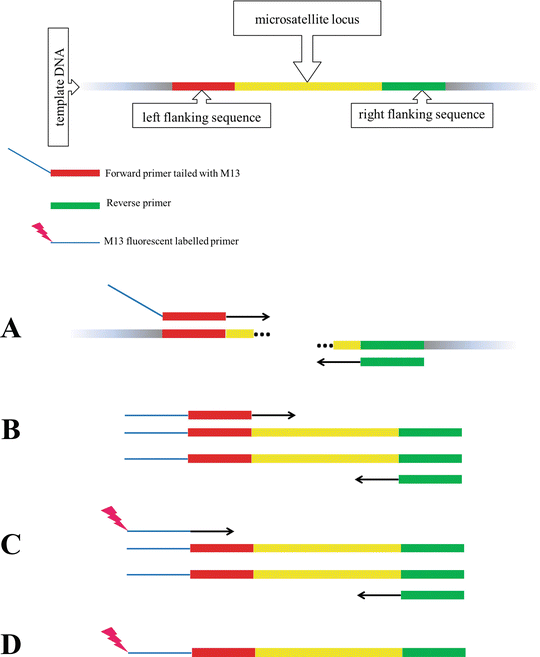
Primer and amplicon construction. The first round of PCR uses a forward... | Download Scientific Diagram